- 1. Principles of Polymerase Chain Reaction Assignment
- 2. Performing a PCR reaction
- 3.(a) Set up the thermocycler
- (b) Explain and Draw the steps
- 4.(a) Double-stranded template of Stand A and Stand B
- (b) Bonds that keep the strand together
- (c) Process Name
- 5.(a) Forward ad reverse Primer sequence for each DNA template strand
- (b) Process of Primer attachment
- (c) Calculate the annealing temperature
- 6.(a) Widely used method of analysis of the PCR
- (b) The complementary strand
- (c) Commonly used enzyme
- 7. Reaction continues for more cycles
- 8. Two different PCR ingredient tests
- 9. Correct PCR product size
- 10. (a) Identify the guilty suspect
- (b) Including the fingerprint of Victim’s DNA
- 11. 3 things about PCR
1. Principles of Polymerase Chain Reaction Assignment
PCR stands for Polymerase Chain Reaction and is a technique used to make more of a specific DNA sequence. It involves particularly three steps which are denaturation that is mainly heating for breaking the overall bonds of hydrogen between the specifically two DNA strands. Annealing which defines the overall binding of the different types of primers to complementary types of sequences within the overall target DNA, along with extension that is the synthesis of particularly new type of DNA strands through the help of Taq polymerase (Zhu et al. 2020). It reoccurs 30-40 times which mainly leads towards a very rapid or strong duplication of the DNA quantity. PCR is specifically used in almost each of the molecular biology laboratories today because of the reasons such as sensitivity, specificity along with ability to get a large quantity of DNA from a little amount of starting material (Kang et al. 2021). It is used in diagnostics, genetic testing, cloning, and in forensic investigation.
Assignment deadlines piling up? Let New Assignment Help ease your burden with expert solutions tailored for student success.
2. Performing a PCR reaction
In order to perform PCR reaction, you simply add some different components into a small and thin tube. These mainly include the overall template DNA that specifically holds the overall sequence to be amplified, along with the primers which are the smaller single stranded DNA sequences that specifically flank the overall target DNA sequence that to be amplified (Olmedillas-López et al. 2022). Deoxynucleotides or dNTPs are also specifically needed for serving as the precursors from which new specific DNA can be synthesized. Taq DNA polymerase, which specifies the heat stable type of enzyme, aids within the synthesis of the DNA during the extension step of the cycle of PCR. This specifically prevents the overall precipitation of the components that should mainly remain dissolved within the solution along with stabilizing the overall activity of Taq polymerase (Kang et al. 2021). The buffer solution also specifically contains magnesium ions or Mg²⁺, while a stabilizer is also specifically added for maintaining the optimal type concentration of enzymes within terms of pH and ionic type strength. These components specifically display a synergistic type of behavior within that they boost the overall DNA sequence.
3.(a) Set up the thermocycler
| STEP | TEMP (°C) | TIME |
| Denaturation | ~94-98 | 20-30 seconds |
| Annealing | ~50-65 | 20-40 seconds |
| Extension | ~72 | 1 minute per kb |
Table 1: Thermocycler Temperature
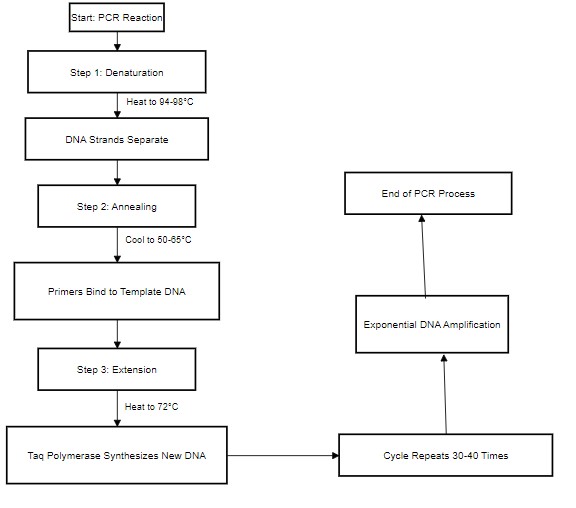
(b) Explain and Draw the steps
Denaturation: The particular DNA of double helix is specifically heated approximately 94 towards 98°C that specifically causes hydrogen type bonds to break and separate the overall strands throughout single type strands.
Annealing: The overall temperature is mainly lowered from approximately 50°C towards 65°C that specifically allows primers to specifically bind towards their complementary type of sequences upon the single-stranded type of DNA (Syamsidi et al. 2021).
Extension: At the temperature of approximately 72°C, Taq polymerase specifically synthesizes a new amount of DNA that mainly strand by adding overall dNTPs complementary towards the particular template strand.
Figure 1: Steps of thermocycler
(Source: Draw.io)
4.(a) Double-stranded template of Stand A and Stand B
Strand A: 5’ GGAAGTGTTTAAAACC...TGGCTGTGATTTCAGTATATG 3’
Strand B: 3’ CCTTCAAAATTTTGG...ACCGACACTAAAGTCATATAC 5’
The strands are antiparallel.
(b) Bonds that keep the strand together
In this process specifically Hydrogen bonds between complementary type base pairs which are A-T: 2 bonds, G-C: 3 bonds.
(c) Process Name
The process is mainly called denaturation.
5.(a) Forward ad reverse Primer sequence for each DNA template strand
Strand A Forward Primer is denoted as GGAAGTGTTT.
Strand B Reverse Primer is denoted as TGGCTGTGAT.
(b) Process of Primer attachment
This particular primer attachment process is mainly called annealing.
(c) Calculate the annealing temperature
Formula is given below.
Tm=2×(A+T)+4×(G+C)
For primers the Forward Primer and Reverse Primer calculations are given below.
- Forward Primer: GGAAGTGTTT
Tm=2×6+4×4=12+16=280C - Reverse Primer: TGGCTGTGAT
Tm=2×4+4×6=8+24=320C
6.(a) Widely used method of analysis of the PCR
The most deeply utilized method of analyzing the products of PCR is Agarose gel electrophoresis.
(b) The complementary strand
The particular name of the complementary type strand is Taq polymerase.
(c) Commonly used enzyme
Taq polymerase is specifically a commonly utilized enzyme because this particular enzyme is totally thermo stable.
7. Reaction continues for more cycles
During PCR reaction when the reaction proceeds for more cycles which defines normally 30-40 cycles, the amount of DNA produced is exponential. Here are the following things which happened during this process.
Exponential Amplification: PCR should increase the quantity of the target DNA fragment after every cycle by two fold (Teymouri et al. 2021). This means that by the end of the first cycle, you have two target sequences, by the second cycle four target sequences, and so forth. After the 30-40 cycle, billions of the target DNA are made and generated for detection and identification.
Plateau Effect: Over time the amplification of the reaction may become less efficient the reaction and may stop altogether once it reaches a certain optimal point in a several cycle reaction. This can happen due to the exhaustion of basic reagents such as primers or nucleotides or presence of inhibitors (Louws et al. 2024). Therefore the specific type of DNA continues to be amplified. The process mainly slows down towards a certain point and reaches its climax.
Non-specific Amplification: As for the specific cycles in succeeding cycles, difficulty is often experienced with non-specific amplification and generation of primer-dimers that are unwanted short products (Madadelahi et al. 2024). To specifically stop this, reaction parameters are mainly including the overall concentration of primers along with cycling numbers that are carefully controlled.
In conclusion PCR cycles means that DNA amplification is exponential but may reach a maximum level of non-specific products. Optimization can be effected to guarantee successful amplification in the required cycle range.
8. Two different PCR ingredient tests
The PCR ingredient that would be different between the two tests for Cystic Fibrosis and Alzheimer’s disease is the Primers.
First, the Primers are short single strand DNA sequences that are required for the start of the amplification of DNA in PCR. They must specifically be able to attach to the sequences of DNA that are on the two sides of the area of interest so that the DNA polymerase can be placed in the correct place to start the synthesis or receipt (Sule and Oluwayelu, 2020). Cystic Fibrosis and Alzheimer’s disease are caused by different gene mutations hence the primers used in that PCR test would be different, they would have different DNA sequences of these diseases.
Defining Cystic Fibrosis, the primers would then be used in a way that enables them to only work on the area, or gene, in the body that causes the Cystic Fibrosis, which is known as the CFTR gene. For Cystic Fibrosis most common form is led by mutation in the CFTR gene which is predominant (Chen et al. 2022). This mutation would be detected by use of primers to successfully pry open the specific region of DNA, where this mutation exists and thus identify any kinds of variations in the Genome sequence that may suggest the presence of the disorder.
In the case of Alzheimer’s disease, especially while examining the genetic risk factors, the primers would be designed in the sections of gene such as APOE or in any other gene related to Alzheimer’s risk. The primers in this case would be to target the genes in or near various genes that could then be screened for specific type mutations or risk alleles.
Other neutral components of the PCR process, the template DNA, nucleotides and Taq polymerase used are similar for the two tests but the primers used differ (Tahamtan and Ardebili, 2020). The choice of primers makes certain that PCR enlarges the unique section of the genome that carries the mutation or risk factor for the given disorder that is being investigated.
Primers are the PCR component that would be different between the test for Cystic Fibrosis and Alzheimer’s disease because primers are synthesized to match specific sequences in the genome related to the disease.
9. Correct PCR product size
Lane 3 shows the right size of the PCR product which is 200 bp. This was done based on the observation of parallel progression of DNA band with that of the 200 bp in the DNA ladder. It is used as molecular weight standards, thus DNA ladders are used to estimate the size of the fragments. The bands in Lanes 1 and 2 are again not corresponding to the 200 bp marker which endorses the result that Lane 3 contains a DNA fragment which was amplified to 200 bp. This alignment also helps in correct identification of the PCR product.
10. (a) Identify the guilty suspect
The guilty suspect is Suspect 2 since his DNA fingerprint matches the DNA specimen from the victim. This conclusion was drawn due to the realization of same matched bands between the specimen and Suspect 2. The bands patterns of the Suspect 1 and Suspect 3 are dissimilar to those of the specimen therefore excluding them. Forensic nucleic acid identification relies on a comparison of specific DNA banding patterns of individual organisms that is multilaterally identical only with monozygotic types of twins (Harshitha and Arunraj, 2021). The matching type of bands between the specimen along with Suspect 2 ascertain that the overall DNA present at the particular crime scene specifically belongs towards this suspect thus mainly proving the complicity of this specific suspect within the incident. This method is rather helpful for a forensic type investigation because of its high amount of specificity.
(b) Including the fingerprint of Victim’s DNA
Distinguishes DNA victim from the DNA perpetrator: All these components are added to the specimen to guarantee that the DNA extracted from it could be indisputably originating from a distinct third party other than the victim’s finger itself.
Quality control: Thus, it helps to find out that the sample was contaminated or the analysis was made in error, and the profile coincides only with the external suspect.
11. 3 things about PCR
As a particular type of result of this activity the user specifically found out about the following facts on PCR or Polymerase Chain Reaction which are given below.
The Primers Specificity within PCR: In the particular reaction, the user specifically understood that only a primer is very essential to control the PCR process. They are first hand simple sequences of DNA that can only attach to the specific region of the target DNA and avoid any non desired fragment to be multiplied. These primers can be designed taking much importance in polymerase chain reaction because they must be specific in such sequences that flank the target region (Tahamtan and Ardebili, 2020). This specificity makes it possible to amplify accurately the required piece of the gene and avoid other non-sense amplification of other sequences.
Thermocycling along with its Phases: From the particular analysis of PCR, the user specifically developed an thermocycling appreciation that specifically applies different types of temperatures upon the specific type of material. The process specifically involves three main steps which are given below. Those specifically involve denaturation, annealing along with extension. Denaturation is specifically performed at avery amount of high temperatures about 94 towards 98°C for producing single stranded type of DNA from the double stranded type of DNA. In the specific cooling phase from 50°C towards 65°C (Louws et al. 2024). The primers stick on to the template DNA during the particular extension step that usually has a temperature of 72° C. Taq polymerase specifically makes a new type of strand of DNA thus mainly amplifying the total amount of DNA within a single amount of cycle. It is very important that one has done their specific type of homework well in as much as dealing with PCR in order to understand these stages.
Application of PCR within Genetic Testing: The user discovered that quantitative PCR is also valuable in finding gene alterations that connect varieties of diseases including Cystic Fibrosis or Alzheimer’s disease. PCR has the ability to enrich segments of the genome of interest such that it can be used for a diagnostic assay to check for mutations or risk factors. PCR comes more into its own in medical diagnosis, research and forensic sciences due to this ability to target specific sequences.
The reader not only acquired a better understanding of PCR and its uses in biotechnology and medical genetics, but also developed an excellent and compact background on the subject.
Reference List
Journals
- Chen, S., Sun, Y., Fan, F., Chen, S., Zhang, Y., Zhang, Y., Meng, X. and Lin, J.M., 2022. Present status of microfluidic PCR chip in nucleic acid detection and future perspective. TrAC Trends in Analytical Chemistry, 157, p.116737.
- Harshitha, R. and Arunraj, D.R., 2021. Real‐time quantitative PCR: A tool for absolute and relative quantification. Biochemistry and Molecular Biology Education, 49(5), pp.800-812.
- Kang, B.H., Lee, Y., Yu, E.S., Na, H., Kang, M., Huh, H.J. and Jeong, K.H., 2021. Ultrafast and real-time nanoplasmonic on-chip polymerase chain reaction for rapid and quantitative molecular diagnostics. ACS nano, 15(6), pp.10194-10202.
- Kang, B.H., Lee, Y., Yu, E.S., Na, H., Kang, M., Huh, H.J. and Jeong, K.H., 2021. Ultrafast and real-time nanoplasmonic on-chip polymerase chain reaction for rapid and quantitative molecular diagnostics. ACS nano, 15(6), pp.10194-10202.
- Louws, F.J., Schneider, M. and de Bruijn, F.J., 2024. Assessing Genetic Diversity of Microbes Using Repetitive SequenceBased PCR (rep-PCR). In Environmental Applications of Nucleic Acid Amplification Technology (pp. 63-94). CRC Press.
- Madadelahi, M., Agarwal, R., Martinez-Chapa, S.O. and Madou, M.J., 2024. A roadmap to high-speed polymerase chain reaction (PCR): COVID-19 as a technology accelerator. Biosensors and Bioelectronics, 246, p.115830.
- Olmedillas-López, S., Olivera-Salazar, R., García-Arranz, M. and García-Olmo, D., 2022. Current and emerging applications of droplet digital PCR in oncology: an updated review. Molecular Diagnosis & Therapy, 26(1), pp.61-87.
- Sule, W.F. and Oluwayelu, D.O., 2020. Real-time RT-PCR for COVID-19 diagnosis: challenges and prospects. The Pan African Medical Journal, 35(Suppl 2).
- Syamsidi, A., Aanisah, N., Fiqram, R. and Al Jultri, I., 2021. Primer design and analysis for detection of mecA gene. Journal of Tropical Pharmacy and Chemistry, 5(3), pp.245-253.
- Tahamtan, A. and Ardebili, A., 2020. Real-time RT-PCR in COVID-19 detection: issues affecting the results. Expert review of molecular diagnostics, 20(5), pp.453-454.
- Teymouri, M., Mollazadeh, S., Mortazavi, H., Ghale-Noie, Z.N., Keyvani, V., Aghababaei, F., Hamblin, M.R., Abbaszadeh-Goudarzi, G., Pourghadamyari, H., Hashemian, S.M.R. and Mirzaei, H., 2021. Recent advances and challenges of RT-PCR tests for the diagnosis of COVID-19. Pathology-Research and Practice, 221, p.153443.
- Zhu, H., Zhang, H., Xu, Y., Laššáková, S., Korabečná, M. and Neužil, P., 2020. PCR past, present and future. Biotechniques, 69(4), pp.317-325. (1 2)